Can you accurately predict how much drug absorbs into the bloodstream?
Drug absorption can be affected by both drug-specific and patient-specific factors. Absorption data determines how much of the drug reaches the systemic circulation after oral or inhaled administration so that a suitable therapeutic range can be established.
Due to their physiological complexity, recreating human-representative biological barriers in vitro is difficult. Animals offer limited advantages as their biological makeup differs from humans. Combined, this means accurately predicting drug absorption and permeability in preclinical drug development can be challenging.
Inhaled medicines offer an advantage over oral administration, due to the low metabolic capacity of the lung. Unfortunately, there are relatively few in vitro inhaled drug absorption in vitro assays available. In vitro gut absorption models suffer from low metabolic capacity, which limits their ability to accurately mimic the human intestinal response to orally administered drugs. Both suffer from permeability values that are not physiologically relevant.
Our solution
PhysioMimix® Drug absorption assays utilize Lung-on-a-chip (alveolar or bronchial), or Gut-on-a-chip co-culture models. Fundamental to the success of these models is organ-specific perfusion, which drives the development of complex 3D tissue morphologies, the expression of key cellular components (e.g., mucus or metabolic enzymes), and barrier integrities that align closely with the human.
These models can be used to calculate absorption into cells and/or drug permeability rates across the epithelial barrier. The oral drug absorption assay can also predict systemic exposure following oral dosing via the first-pass metabolism. Data derived using these assays more accurately translates into human outcomes compared to traditional approaches.
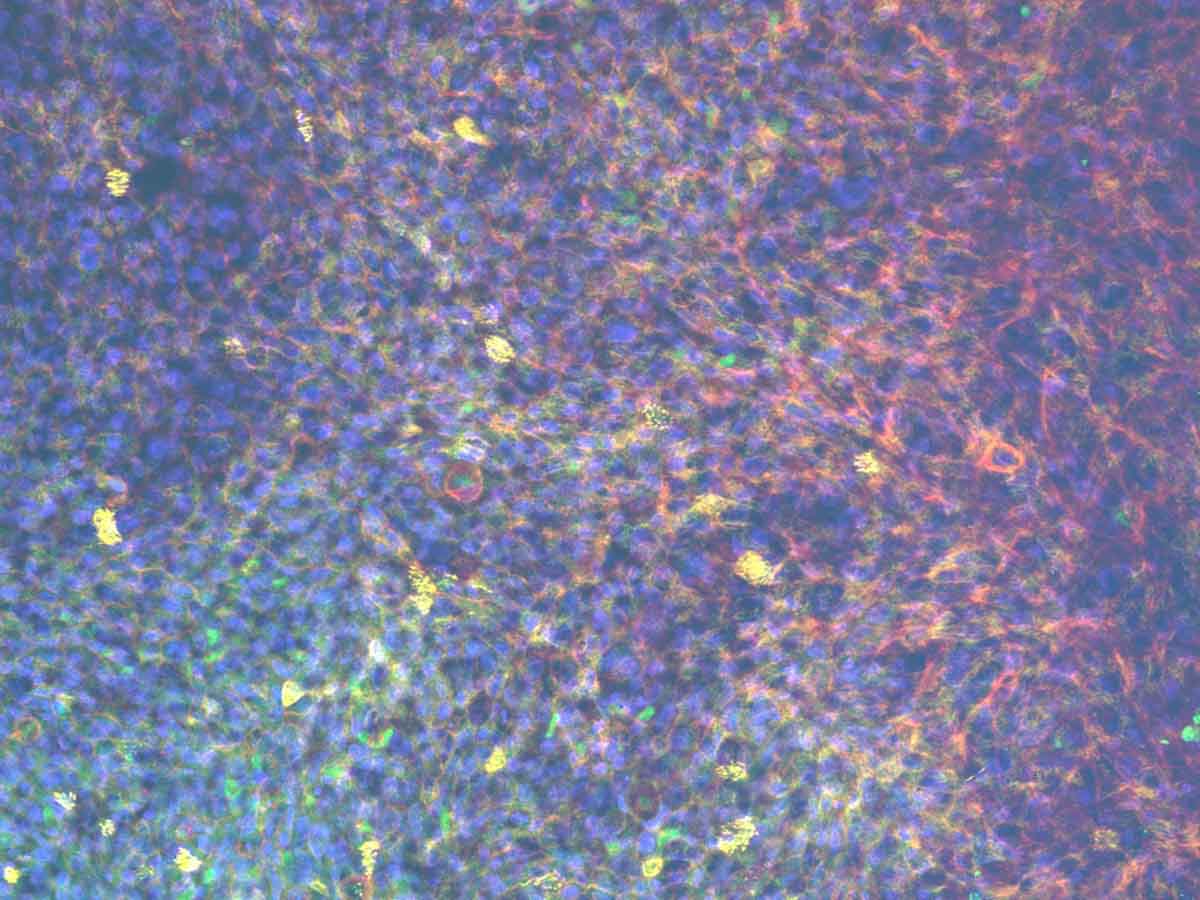
Studying drug absorption
Limitations with current techniques
- Difficult to predict transport rates for slowly absorbed compounds
- Lack of mucous secretion
- Limited transporter expression/activity
- Low metabolic activity
- In vivo model data fail to translate into clinical studies
Advancements with PhysioMimix Core
- Perfusion of microtissues increases in-vivo-like barrier integrity and permeability of the epithelium
- Goblet cells secrete mucus, more closely mimicking the mucosal barrier
- Major transporters are expressed by the epithelium
- Organ-relevant metabolic capacity
- Specific tissue functions and assay types are enabled by the 3D complexity of the model
End point measurements
Liver and gut longitudinal and endpoint measurements include (but not limited to):
Functionality biomarkers
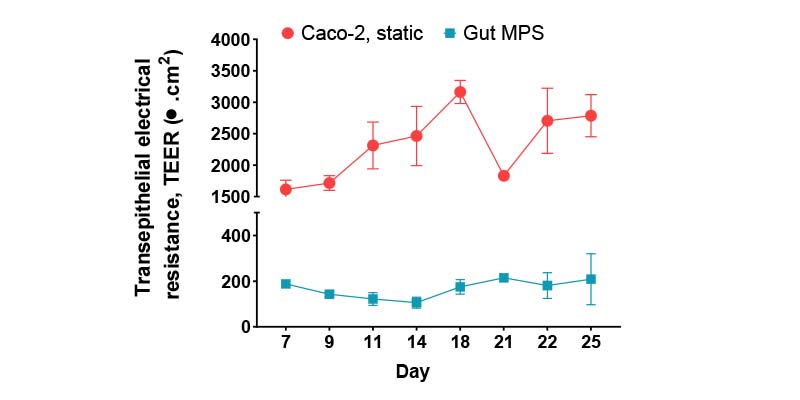
- Trans epithelial electrical resistance (TEER)
- Lactose dehydrogenase (LDH) release
- Lucifer yellow or dextran permeability assay
Profiling analysis
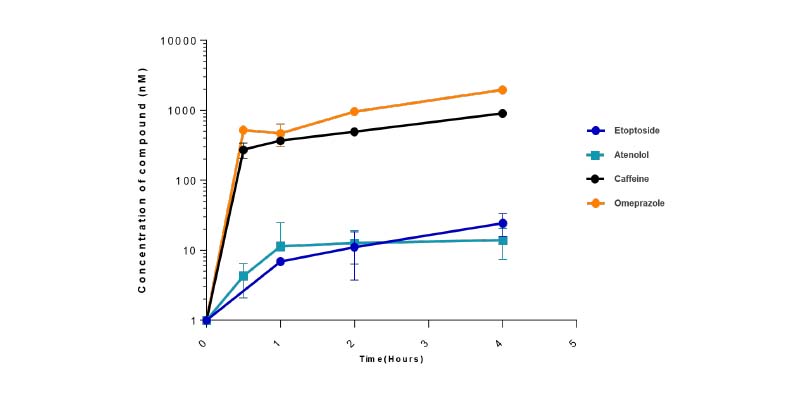
- Media and tissue samples ready for LC/MS analysis
- Phenotypic analysis confirmed by qPCR or microscopy
- Histology, or colorimetric assays to confirm mucus production
Optional profiling analysis
- Quantitative PCR
- Transcriptomics
Explore our Lung-on-a-chip models
Recreate the airway or alveolar niche by culturing epithelial and endothelial cells on each side of an insert at ALI. Achieve a human-relevant barrier with enhanced functionality for longer term culture.
Access our ADME service
Get instant access to PhysioMimix Drug Absorption Assays via our CRO Service. Through a collaborative approach, our experts work with you to plan and execute your study.
Bespoke projects are carried out by our dedicated team of scientists in our CRO facility providing you with actionable data within weeks.
Add PhysioMimix Core to your lab
Harness the power of PhysioMimix Core in your own lab.
With a growing community of users and support from our experts, there has never been a better time to transition into 3D cell culture.