Resource > Articles >
A Guide to Choosing the Right In Vitro Liver Models for Drug Development Tasks
How New Approach Methodologies and Organ-on-a-Chip Technology Are Redefining Drug Development
Filed under: ADME, DILI, Drug bioavailability, Drug metabolism, and Safety toxicology

As the major drug metabolizing organ, the liver is central to safety and DMPK assessments. There are often challenges in accurately predicting human-relevant parameters, particularly as in vivo models largely have wide ranging metabolic profiles compared to human.
However, a diverse set of in vitro liver models are available for use, from simple suspension cultures to advanced microphysiological systems (MPS). No single model is universally best as each provides different strengths, limitations and practical considerations. So, how do you choose the right in vitro liver models for drug development? This article explores why it is imperative to match the model to the specific scientific question you need to answer when selecting the right tool for the job.
How to choose the right in vitro liver models for drug development and discovery
Across our industry, the same challenges continue to hinder decision-making:
- unclear or conflicting animal data
- risk of late stage failures
- need for more mechanistic clarity
Addressing these issues requires looking beyond historical default choices. By utilizing an evidence-based framework, that aligns model capabilities to what you need to predict, detect, or understand, you can reduce the risk of wasted cost and resources. This article explores three common scenarios where liver models diverge in suitability and highlights where MPS provides the clarity to move forward with greater confidence.
Scenario 1a. Predicting drug clearance using in vitro liver models
When the question is – What metabolism affects my molecules and how quickly are they cleared? – your model needs to demonstrate:
- high metabolic fidelity
- broad enzyme coverage
- stability over time
An interesting study by Kratochwil et al., (2017) compared the key metabolic activities of the cell line HepG2, iPSC-derived hepatocytes (iCell®), the hepatic stem cell line HepaRG™, and human hepatocyte co-cultures (HμREL™ and HepatoPac®) to freshly isolated primary hepatocyte suspension cultures across 11 enzyme markers. The results identified that the metabolic activity of HepaRG, HepatoPac, and HµREL was broadly comparable to PHH suspensions, with some CYP2D6 and non-CYP differences. By contrast, iPSC-derived hepatocytes and HepG2 cells exhibited tenfold lower metabolic activity and were therefore identified as less suited for clearance predictions or metabolism-driven assessments.
HepaRGs offer advantages in cost and scalability but can underpredict due to slower metabolic turnover, whilst metabolic activity in PHH suspension assays is short-lived (hours). To extend study windows, researchers in the past have turned to HµREL (routinely run over 72 hours) or HepatoPac (routinely up to seven days) formats, which offer a more reliable yet practical approach for profiling.
However, certain questions require longer experimental windows, including clearance studies beyond a week, or studying enzyme induction – which typically requires multiple dosing cycles. Additionally, more sophisticated systems may be required to assess the combined effects of metabolism by the gut and liver to improve oral bioavailability predictions.
Whilst currently impractical for high-throughput work, complementary perfused Liver MPS (also known as Liver-on-a-chip) overcome the limitations of standard tools where the highest possible metabolic relevance is required. With either 12- or 48- individually perfused liver chips per MPS plate, the dynamic flow conditions of Liver MPS extend metabolic performance over multiple weeks and enables individual organ models, such as human gut and liver, to be interconnected to emulate the dynamics of drug absorption through the intestinal barrier, followed by their subsequent metabolism by the liver. This innovative in vitro approach allows intravenous and oral dosing to be compared for the prediction of human oral bioavailability (Abbas et al., 2025). By answering these previously unaddressable questions, insights from MPS better inform clinical study design.
Scenario 1b. Predictive models for metabolite-induced DILI assessment
Taking this a step further, where your question relates to predicting metabolite-induced DILI, the first consideration is whether a simple yes/no triage is sufficient, or do you need deeper mechanistic insights?
Spheroid models reporting macro changes to ATP levels (indicating cell death) represent a practical early screening tool to improve human relevance over 2D assays, but their sensitivity is often restricted by metabolic capacity and longevity. Rubiano et al., (2020), demonstrated that PHH CYP3A4 activity was significantly higher and more stable (up to 28 days) in PhysioMimix Liver MPS cultures compared to spheroids or sandwich cultures. Mechanistic confirmation via pathway interrogation and the identification of latent hepatotoxicity also benefit from the enhanced capabilities of a fully perfused Liver MPS model comprised of PHHs and Kupffer cells.
This is particularly relevant for idiosyncratic DILI and metabolite-dependent bile acid transporter interference, both of which can develop slowly as metabolites accumulate. In the same study by Rubiano et al. (2020), it was demonstrated that the Liver MPS was able to capture both Phase I and II metabolism of troglitazone to its toxic metabolites. Phase II metabolism can often differ between species, therefore a human-relevant model is important to capture these events. Combining the use of spheroids for early screening and MPS for confirmation and mechanistic evaluation therefore delivers a more comprehensive workflow for reducing late-stage risk.
Scenario 2. Detecting cholestasis mechanisms in vitro
When the question becomes “do my lead candidates pose cholestasis risks?” and “can this model detect the mechanistic signatures of cholestasis”, the requirements shift. Essential are:
- high bile acid transport and metabolism fidelity
- stable long-term function,
- a panel of cholestasis-relevant endpoints
MPS systems are the primary option, but the ability and sensitivity to detect cholestasis varies. In a comparative study, Nitsche et al., (2025) highlighted that, when compared across platforms, only CN Bio’s PhysioMimix Liver MPS detected consistent bile acid reductions and early mechanistic changes across cholestatic toxicants, including bosentan and α-naphthyl isocyanate. Both perfused HepaRG and PHH cultures reported changes in the total bile acid content as a late-stage biomarker of cholestatic DILI -but with different conjugation profiles. Glycine-conjugated bile acids were found to be most abundant in PHH cultures, consistent with human liver, while HepaRG mainly produced taurine conjugates.
This suggests an optimal, staged workflow consisting of:
- Perfused HepaRG Liver MPS using PhysioMimix Core for initial cost-efficient risk identification
- Perfused PHH Liver MPS using PhysioMimix Core for mechanistic confirmation with clinically aligned biomarkers
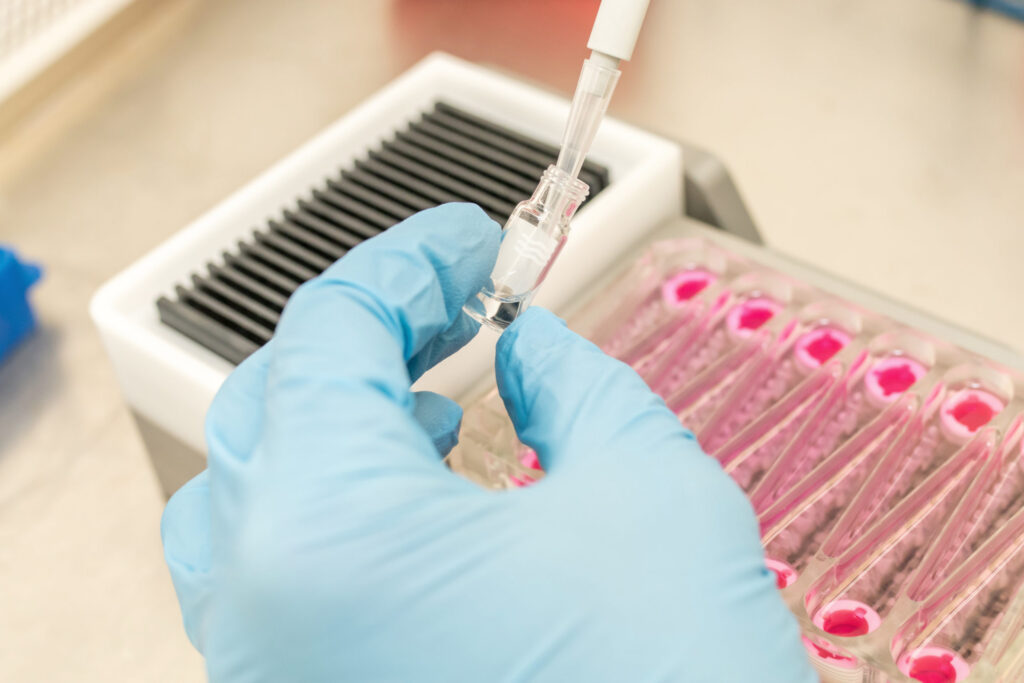
Using PhysioMimix Core’s Liver MPS, bile acids are measured in media (via kits or LC-MS methods) over the dosing period, whilst bile acid synthesis, metabolism and transporter expression can be explored via OMICs analysis of recovered microtissues at the end of a study.
Scenario 3. Understand if preclinical in vivo toxicity will translate to humans with cross-species hepatotoxicity translation models
A third challenge is answering “which animal species predicts the human response”, when two animal species (rodent, non-rodent) give opposing or inconsistent results. This requires cross-species equivalence:
- consistent culture conditions and stable function over >10 days
- ability to detect interspecies divergence
- clinically relevant biomarker reporting
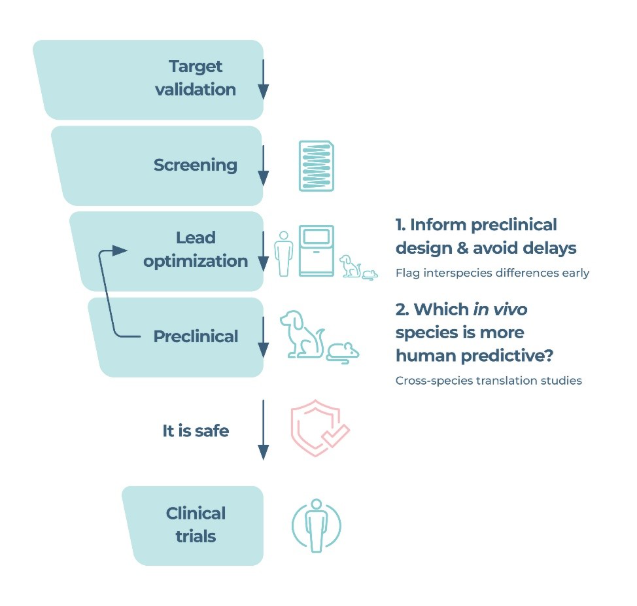
In 2025, Negi et al., compared primary hepatocyte function and drug effects in human, monkey, dog, and rat using a PhysioMimix Liver MPS and 2D assays. MPS assays maintained longer-term function and captured species-specific toxicity responses that 2D systems could not. Whilst MPS is more expensive and lower throughput, the number of drugs to be screened in this context is nominal. Furthermore, a large range of assessments were made, from standard DILI biomarkers (LDH, ALT, AST, albumin, urea) over the dosing period, to transcriptomics of the tissues at endpoint to determine species differences in DILI response at the molecular level. By comparing rodent and non-rodent outcomes to the human MPS, further confidence can be given to either progressing candidates or supporting no-go decisions.
Further to the application of cross-species MPS in troubleshooting inconsistent results from two-animal species safety testing, MPS can also be used pre-emptively to predict species applicability for safety testing or flag any interspecies differences early. Insights from these assays can therefore mitigate the risk of late-stage conflicting data, or save safe candidates being wrongly abandoned or toxic candidates being mistakenly classified as safe. These in vitro tests can also be used to reduce the number of animals being tested in preclinical safety testing, as well as safeguarding animals from exposure to potentially toxic agents, thus supporting 3Rs objectives and ethical considerations. Together, , it can be argued that the translational clarity offered by MPS to minimize the risk of drug misclassification, support more confident clinical development progression, and prevent unnecessary loss of safe assets from the pipeline is justified.
Summary
Rather than asking which model is ‘best,’ or relying on historical decisions – ask which model is best for the question you need to answer at the particular drug discovery or development phase you operate. Each liver model provides different strengths in metabolic competence, mechanistic capability, data translatability, practicality and cost. By understanding and strategically combining these tools, you can derisk programmes earlier with human-relevant data that reduces reliance on animal studies.
As global regulatory momentum shifts towards NAMs approaches, modernizing workflows is now a must, and MPS are no longer “nice to have” technology. They are a risk reduction necessity for guiding decision making and building holistic, mechanistically informed regulatory packages that accelerate the path to market. So, the next time you ask yourself this question, consider MPS.
This article was originally published in a shorter form in the Spring Edition of Eurolab Magazine and on ScientistLive and has been re-published with the kind permission. A link to the orignal version.

Dr. Emily Richardson
Biology Group Leader
Her work bridges complex cell biology and real-world drug discovery, informed by a PhD from the University of Leicester in 3D cell culture for metastatic lung cancer and a BSc in Biochemistry and Molecular Medicine degree from the University of Nottingham.
References:
Find out more about how PhysioMimix® Core de-risks decision-making by generating human mechanistic insight in ADME, Safety Toxicology & Disease modeling applications.
Further reading: Bridging the translational gap with human-relevant ADMET: How New Approach Methodologies and Organ-on-a-Chip Technology Are Redefining Drug Development


